Haemolysin, Catalase and Hydrogen Sulphide Production as Unique Phenotypic Virulence Determinants, Biofilm Formation and Defense against Antibiotics among Mycoplasma arginini and Mycoplasma ovipneumoniae Isolated from Pneumonic Lungs of Domestic Sheep (Ovis aries)
Haemolysin, Catalase and Hydrogen Sulphide Production as Unique Phenotypic Virulence Determinants, Biofilm Formation and Defense against Antibiotics among Mycoplasma arginini and Mycoplasma ovipneumoniae Isolated from Pneumonic Lungs of Domestic Sheep (Ovis aries)
Mona M. Osman1, Kamelia M. Osman2, Manal Abu Elmakarem Mohamed1, Mahmoud E. Hashad2, Jeeser Alves Almeida3, Alaa Saad4, Octavio Luiz Franco5,6, Heba N. Deif 2*
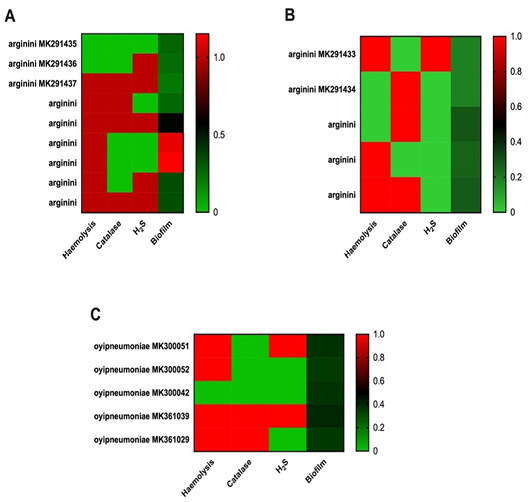
Clustering heatmap of virulence traits phenotypes by lungs and nasal swab.
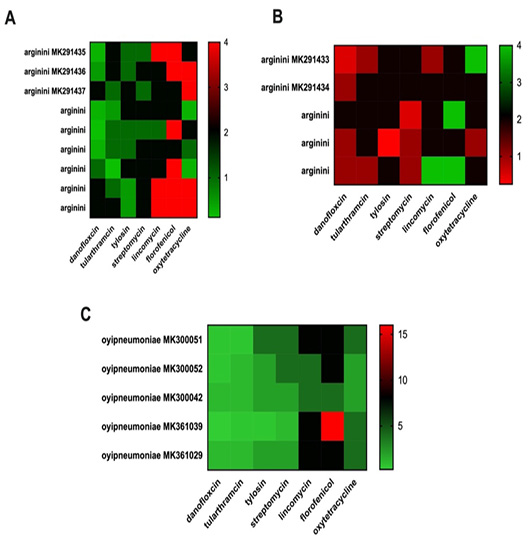
Clustering heatmap of antimicrobial resistance by lungs and nasal swab.
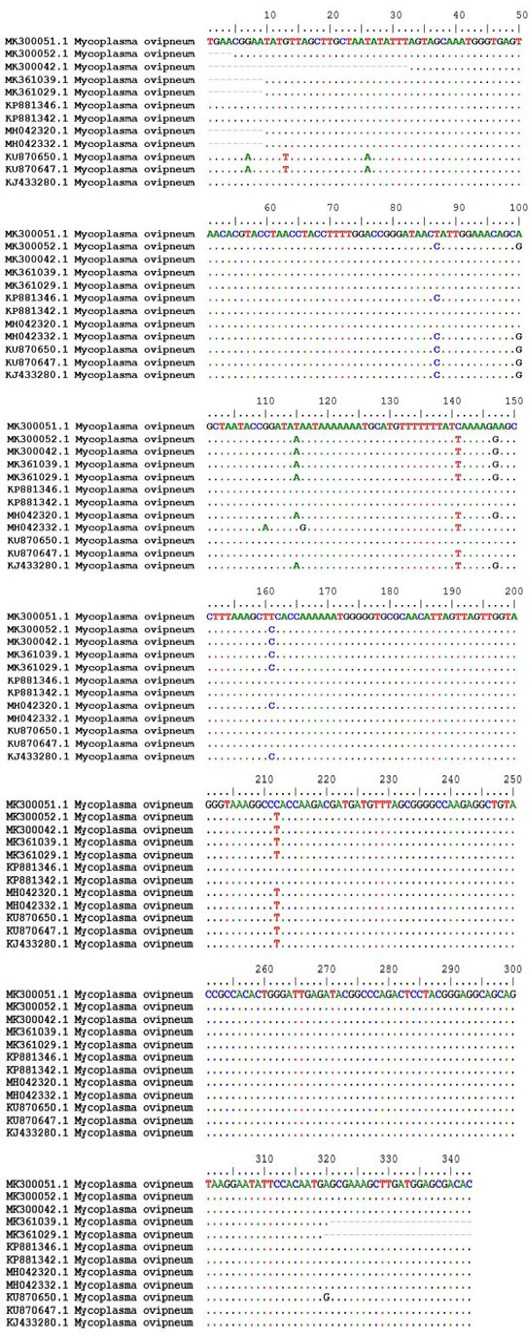
Maximum Likelihood phylogenetic tree of 16S ribosomal RNA gene, partial sequence of sheep Mycoplasma arginini (five isolates) and M. ovipneumoniae (four isolates) in comparison with other representative strains of M. arginini and M. ovipneumoniae available on NCBI.
Maximum Likelihood phylogenetic tree of 16S ribosomal RNA gene, partial sequence of goat M. arginini (four strains) and sheep M. arginini (five strains), Cattle M. arginini (one strain), Camel M. arginini (four strains) in comparison with human M. arginini (two strains) on Genbank, and other representative strains of M. arginini available on NCBI. Camel ━, Sheep ●, Goat □, Human ━ and cattle □ isolates.
Sequence results submitted to the Genbank.