Homozygous Nonsense Mutation in the ASPM Gene Causes MCPH in Consanguineous Pakistani Families
Homozygous Nonsense Mutation in the ASPM Gene Causes MCPH in Consanguineous Pakistani Families
Ansar Ahmed Abbasi1,*, Kathrin Blasius2-4, Imtiaz Ahmed6, Hao Hu7, Sylvie Picker-Minh2-4,8, Muhammad Nasim Khan5, Khalid Hameed1, Aneela Gulnaz1, Zahid Latif5, Abdul Rauf5 and Angela M. Kaindl2-4,8
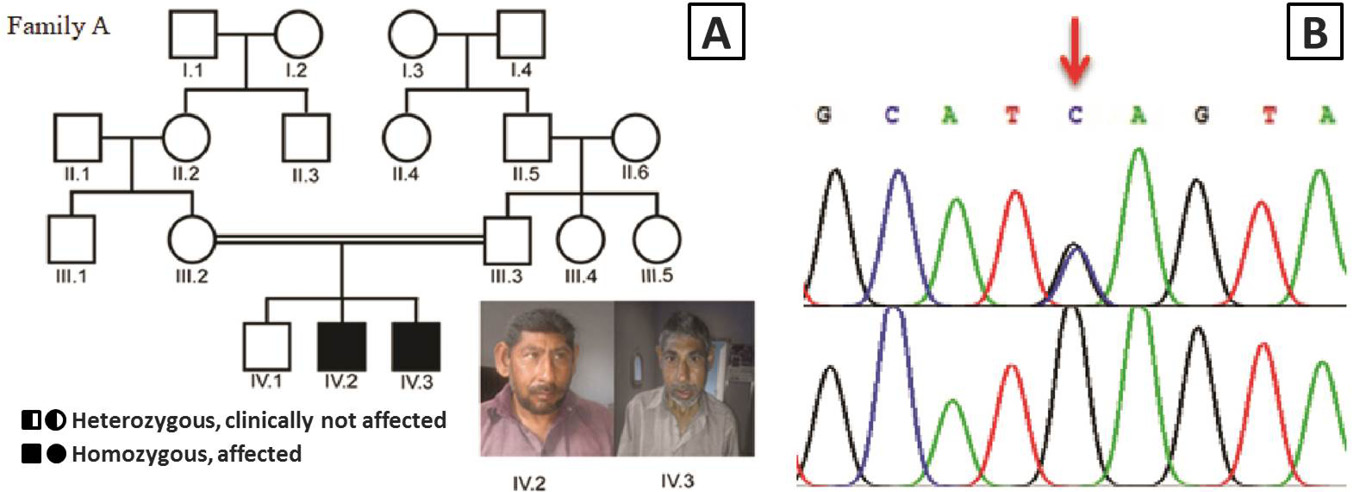
A, Pedigree of the consanguineous family A of Pakistani descent with two patients affected by microcephaly (□, male; ○, female; ═, consanguineous marriage); B, Sanger sequencing results of patient IV.2 from family A showing a homozygous mutation c.4802C>G in the ASPM gene that leads to a nonsense mutation p.S1601* (red arrow). The parental control III.2 is heterozygous for the ASPM mutation.
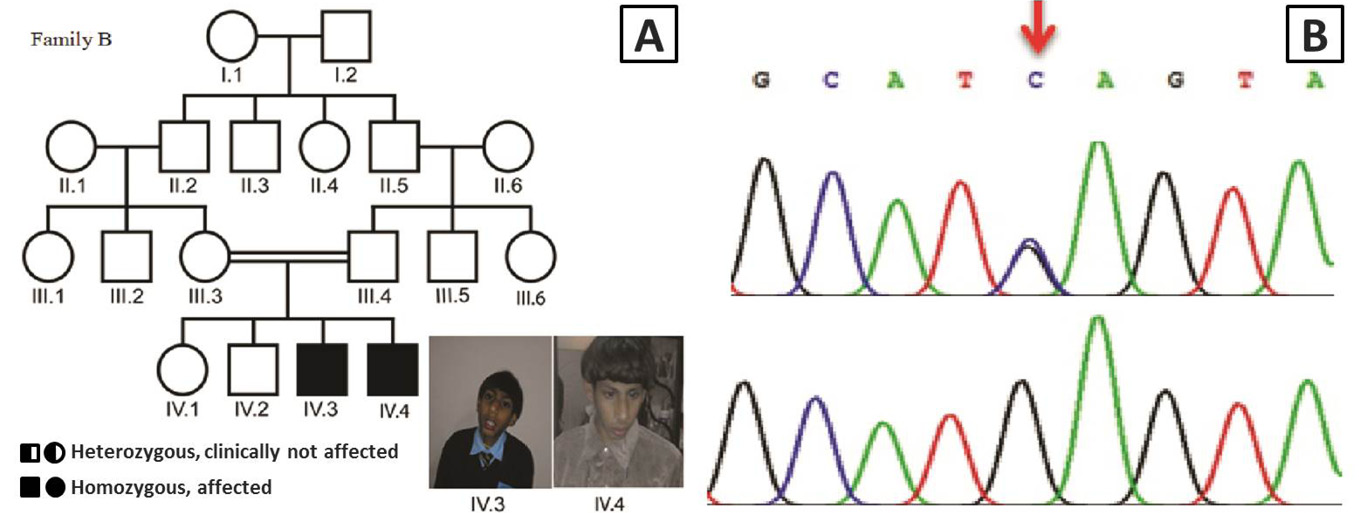
A, Pedigree of the consanguineous family B of Pakistani descent with two patients affected by microcephaly, speech delay and motor delay (□, male; ○, female; ═, consanguineous marriage); B, Sanger sequencing results of patient IV.2 from family B showing a homozygous mutation c.4802C>G in the ASPM gene that leads to a nonsense mutation p.S1601* (red arrow). The parental control III.3 is heterozygous for the ASPM mutation.
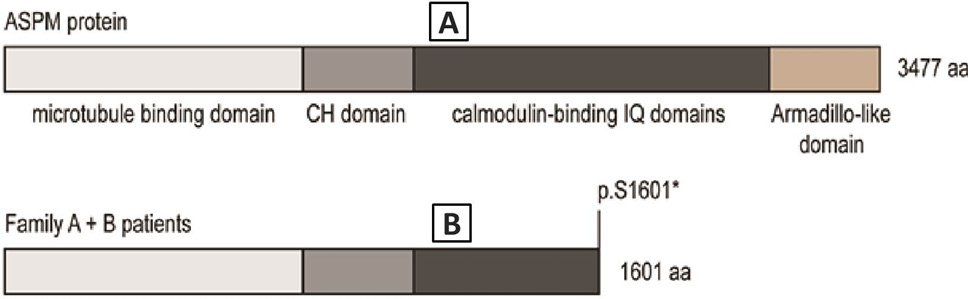
A, the ASPM gene codes for a 3477 aa protein with four protein domains: microtubule binding domain (light grey), CH domain (grey), calmodulin-binding IQ domains (dark grey) and the Armadillo-like domain (brown); B, the nonsense mutation p.S1601* in patients of family A and B lead to a truncated protein, which is missing the Armadillo-like domain and part of the calmodulin-binding IQ domain.