mir-16-5p as a Suitable Reference Gene for Normalization of Quantitative Real Time PCR in Acute Lymphoblastic Leukemia
mir-16-5p as a Suitable Reference Gene for Normalization of Quantitative Real Time PCR in Acute Lymphoblastic Leukemia
Samiah Shahid1,Jawaria Shaheen1, Wajeehah Shahid2, M. Waheed Akhtar1,3 and Saima Sadaf1,*
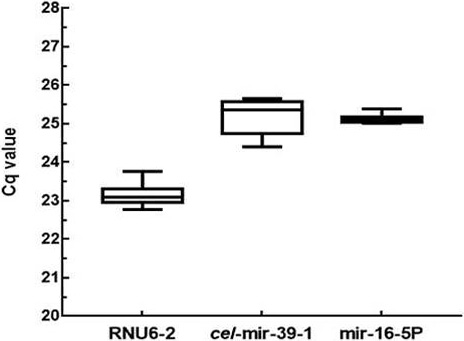
Comparison of quantitative real time PCR for RNU6-2, cel-mir-39-1 and mir-16-5p as reference genes determined through qPCR. Boxes represent mean and standard deviation and whiskers represent minimum and maximum values.
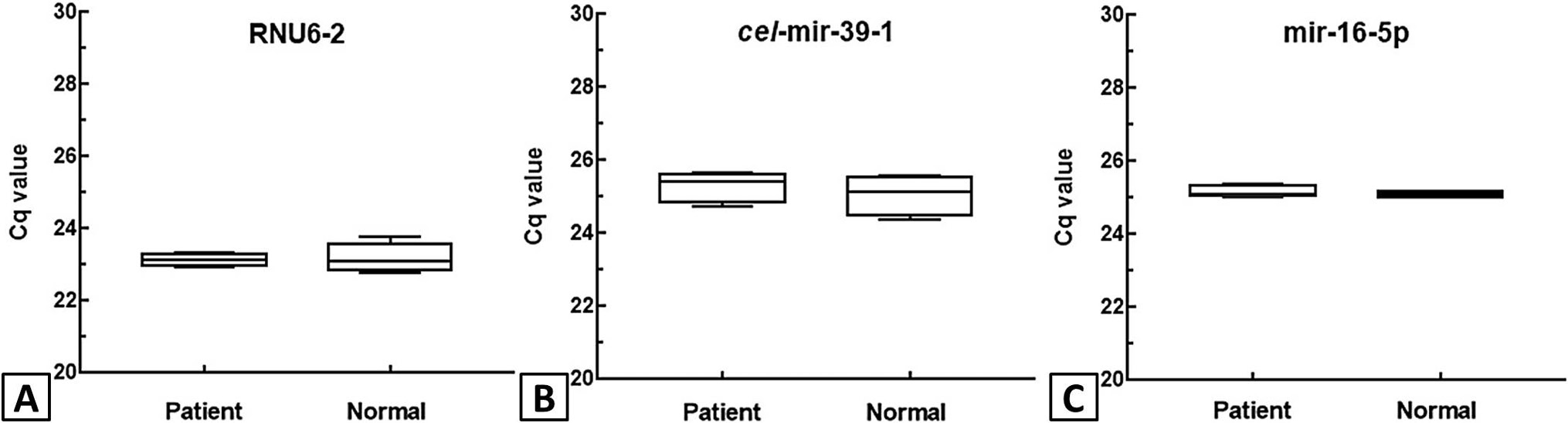
Determination of quantification cycle (Cq value) of the three candidate reference genes RNU6-2 (A) cel-mir-39-1 (B) and mir-16-5p (C) in the patient and normal control samples of ALL determined through qPCR. The values for each distribution are given as mean ± standard deviation as shown in box and whiskers plot.
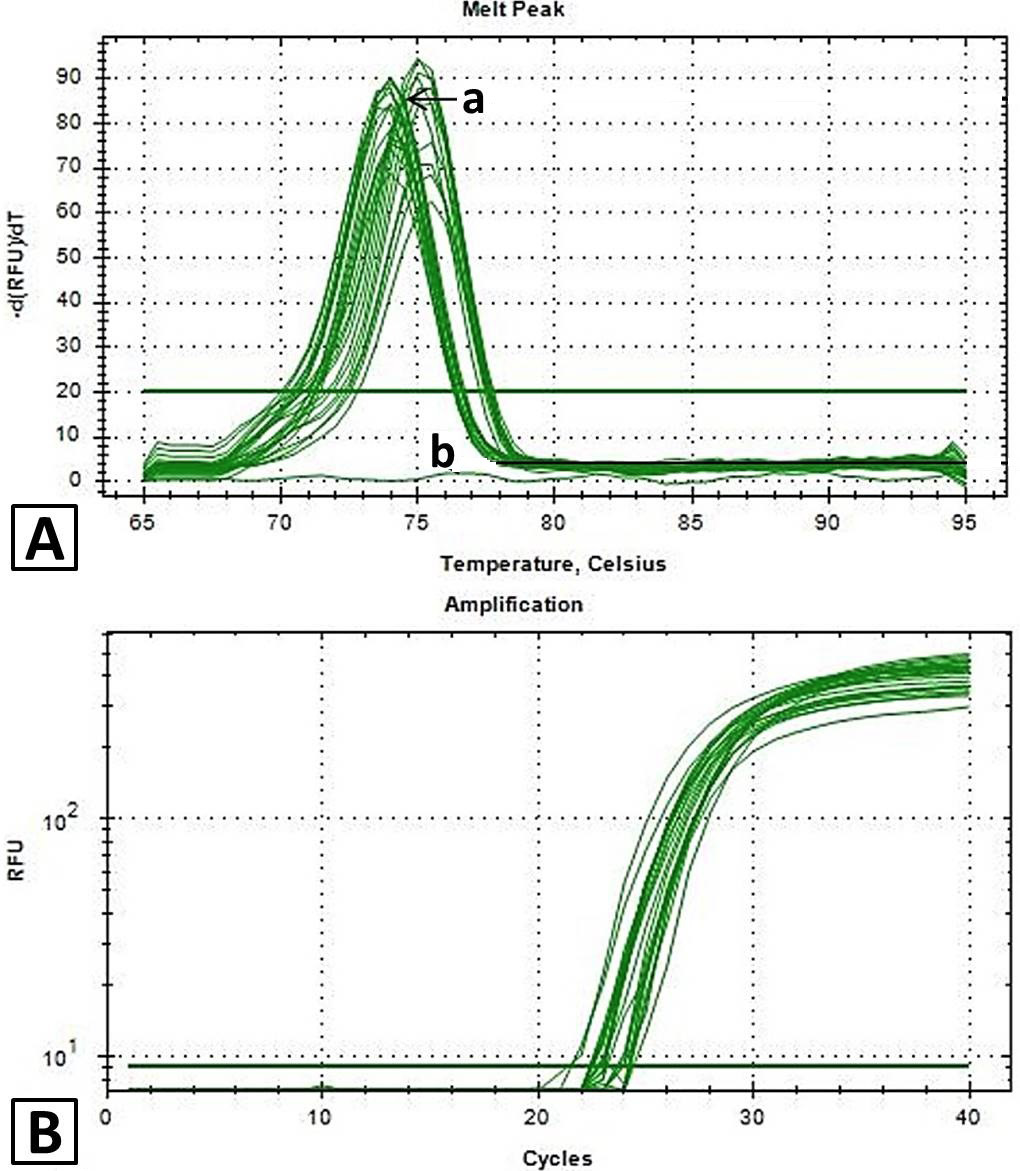
A, melt curve analysis of candidate reference genes for normalization of qPCR in ALL. A singe peak at melting temperature 75.5°C showed amplified products are specific and no non-specific amplification were occurred (a, specific amplification curves; b, NTC-no template control); B, amplification curves of candidate reference genes through qPCR in ALL showing relative fluorescence unit (RFU) of more than 103.
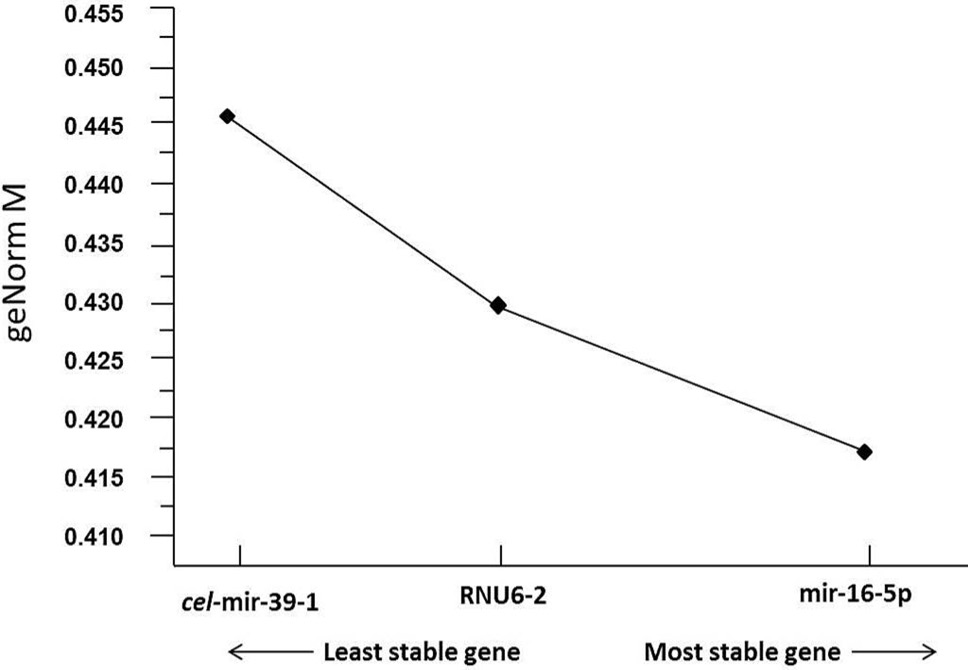
GeNorm analysis of average expression stability by qBase+ software: GeNorm results (M values) for candidate reference genes analyzed (normalized against c. elegans mir-39 as spike in control) indicated mir-16-5p as the most suitable reference gene with highest expression stability.