A Novel ECEL1 Variant Associated with a Congenital Contracture Disorder
Humera Manzoor1,2,5 Norbert Brüggemann2,3, Hafiz Muhammad Jafar Hussain1, Tobias Bäumer2, Frauke Hinrichs2, Muhammad Wajid4, Alexander Münchau2, Katja Lohmann2* and Sadaf Naz1*
1School of Biological Sciences, University of the Punjab, Quaid-i-Azam Campus, Lahore 54590, Pakistan
2Institute of Neurogenetics, University of Luebeck 23538, Lübeck, Germany
3Department of Neurology, University of Luebeck 23538, Lübeck, Germany
4University of Okara, Okara, Pakistan
5Department of Human Genetics and Molecular Biology, University of Health Sciences, Lahore, Pakistan.
* Corresponding author: katja.lohmann@neuro.uni-luebeck.de; naz.sbs@pu.edu.pk
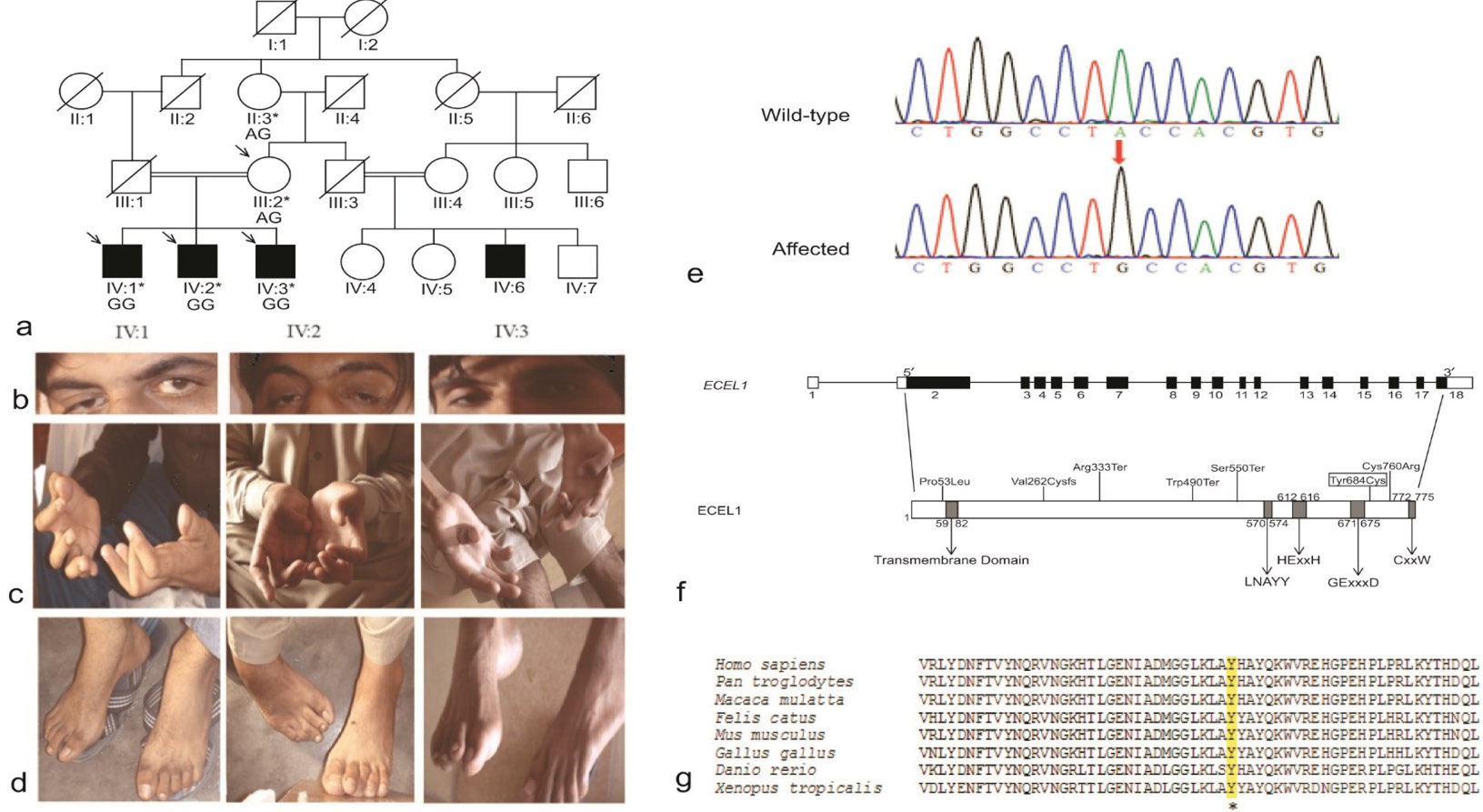
Fig. 1.
Pedigree of family RDHR-01 and genotypes for the ECEL1 variant segregating with the phenotype.
(a) RDHR-01 pedigree. *Individuals who participated in this study. Arrows denote individuals for whom whole-exome sequencing was performed. The genotypes for ECEL1 (c. 2051A>G) p.(Tyr684Cys) variant are indicated below the individual symbols of all participants. Clinical images of affected individuals IV:1, IV:2 and IV:3. (b) Unilateral ptosis severe on right side in individual IV:1 and mild bilateral ptosis in others. (c) curved fingers and adducted thumb of hands. (d) mild camptodactyly of toes in individual IV:1, clubfoot deformity in individual IV:2 and high arched of feet in individual IV:3. (e) Electropherogram of ECEL1 sequence analyses. The site of mutation is depicted by an arrow. (f) Schematic representation of ECEL1 (NM_004826.2), black boxes represent translated exons and plain boxes denote 5ʹ and 3ʹ untranslated regions. Introns are depicted by horizontal lines. The variant p. Tyr684Cys identified in family RDHR-01 is boxed. LNAYY motif is involved in orientation of substrate peptide bond. HExxH is a zinc biding motif. GExxxD is a Zinc coordinating motif. CxxW is the conserved carboxy terminal sequence of metalloprotease. (g) Conservation of ECEL1 residue p. Tyr684 from eight species of vertebrates. The residue p. Tyr684 is highlighted and marked with an asterisk. (Coloured figure may be observed in the online issue of the journal).
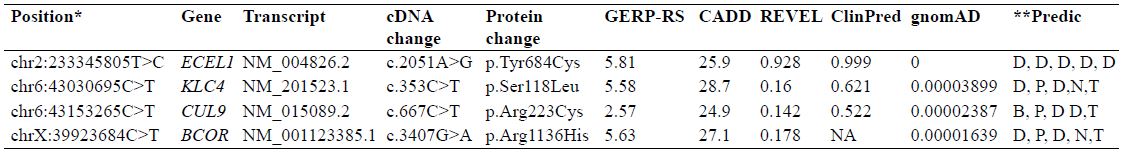
Table II.
Segregating variants in family RDHR-01.
*, position according to the human genome build GRCh37/hg19; GERP-RS, genomic evolutionary rate profiling-rejected substitutions; CADD, combined annotation dependent depletion; REVEL, rare exome variant ensemble learner; ClinPred, (https://sites.google.com/site/clinpred/); ExAC, exome aggregation consortium; ,**, predictions according to PolyPhen2; MutationTaster, PROVEAN, SIFT and FATHMM, respectively; D, deleterious or damaging, P, pathogenic; N, neutral; T, tolerated; B, benign; NA, not available.